Out of 10 and 16 Type II R-M systems detected in the two best-studied H. pylori is unusually rich in restriction-modification (R-M) systems, which generally consist of a methyltransferase (MTase) activity targeting a distinct nucleotide sequence motif, and a cognate restriction endonuclease (REase) that can cleave the same motif when unmethylated. In vitro, mean import lengths depend on donor–recipient combinations, and range from 1.3 to 3.8 kb (refs 21, 22), whereas shorter imports were calculated from comparative sequence analyses of sequential patient isolates (estimated mean lengths: ∼400 bp) 6, 9, 10. Within the cytoplasm, ssDNA interacts with DprA and RecA prior to recombination into the recipient’s genome 19, 20. Double-stranded DNA (dsDNA) is imported into the periplasm via the ComB transport system 16, 17, which is thought to be followed by conversion of dsDNA to single-stranded DNA (ssDNA) and subsequent transfer into the cytoplasm 18. pylori is naturally competent for DNA uptake. pylori, and introduces up to 109 times more substitutions than mutation 14. Recombination has been shown to be the dominant driving force of genetic diversification of H. pylori and leads to the exchange of large proportions of its genome during mixed infections with different strains within one stomach 6, 7, 8, 9, 10, 11, 12, 13, 14, 15. An exceptionally high recombination rate is an additional hallmark of H. The species displays exceptional genetic diversity and variability, partly due to a very high mutation rate 2 that results from a lack of genes encoding a classical methyl-directed mismatch repair (MMR) system ( mutHLS1) 3, 4 and from translesion-synthesis properties of its DNA polymerase I (ref. pylori infection causes chronic active gastritis which can progress to severe diseases, including gastroduodenal ulcers and gastric cancer 1. H elicobacter pylori is one of the most successful bacterial pathogens, infecting the stomachs of more than half of all humans. pylori gene content and its highly recombinational population structure. We conclude that restriction-modification systems inhibit the genomic integration of novel sequences, while they pose no barrier to homeologous recombination, which reconciles the observed stability of the H. In contrast, REases limit the import of heterologous DNA. Restriction endonucleases (REases) of the recipient bacteria fail to inhibit integration of homeologous DNA, independently of methylation. 
pylori genomes from chronically infected people demonstrates the same bimodal import pattern in vivo. Import lengths show a bimodal distribution with averages of 28 and 1,645 bp. A single transformation cycle results in up to 21 imports, and repeated transformations generate a maximum of 92 imports (8% sequence replacement). Here we use an in vitro transformation system combined with genome sequencing to study chromosomal integration patterns after natural transformation. However, please notice that only it is only possible to select one primer at the time for analyzing primer properties.Recombination plays a dominant role in the evolution of the bacterial pathogen Helicobacter pylori, but its dynamics remain incompletely understood. individual primers.Įach primer can now be analyzed for its properties. Your primer sequences will now extracted as single sequences (Figure 1).įigure 1: The sequence in the sequence list are here extracted to single sequences, i.e. Choose to Save your results and finish the wizard.In the next step of the wizard chose Extract to single sequences and click Next.In the wizard that opens up chose your imported sequence list and click Next.Genomics Workbench: Toolbox | Classical Sequence Analysis | General Sequence Analysis | Extract Sequences Main Workbench: Toolbox | General Sequence Analysis | Extract Sequences To extract the primers (sequences) as individual primers go to:.The primers are now imported as a sequence list. csv file into your CLC Workbench using the standard import option and leaving the import setting as automatic. This can be done by selecting CSV (comma delimited) (*.csv) under Save as type. There must not be any empty rows between the primers. The first column should be the primer name and the second column should be the primer sequence.Excel. The Workbench support IUPAC codes for nucleotides as list on the following manual page: IUPAC codes for nucleotides 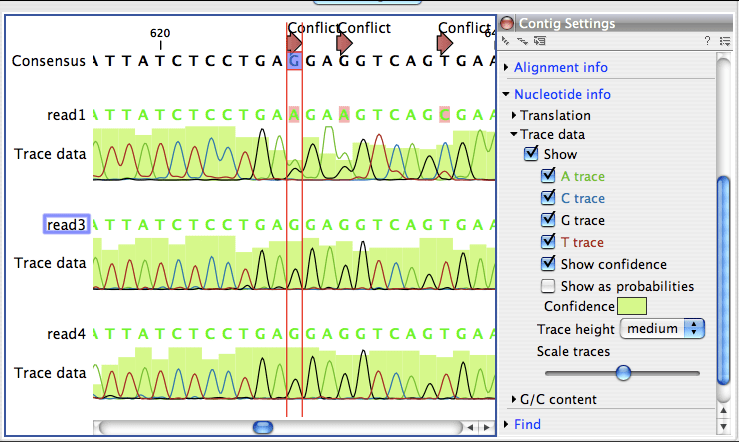
Make a list of the primers that you want to import in e.g.To import primer sequences to a CLC Workbench please follow the steps outlined below: 
How to import predesigned primers to a CLC Workbench?
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
